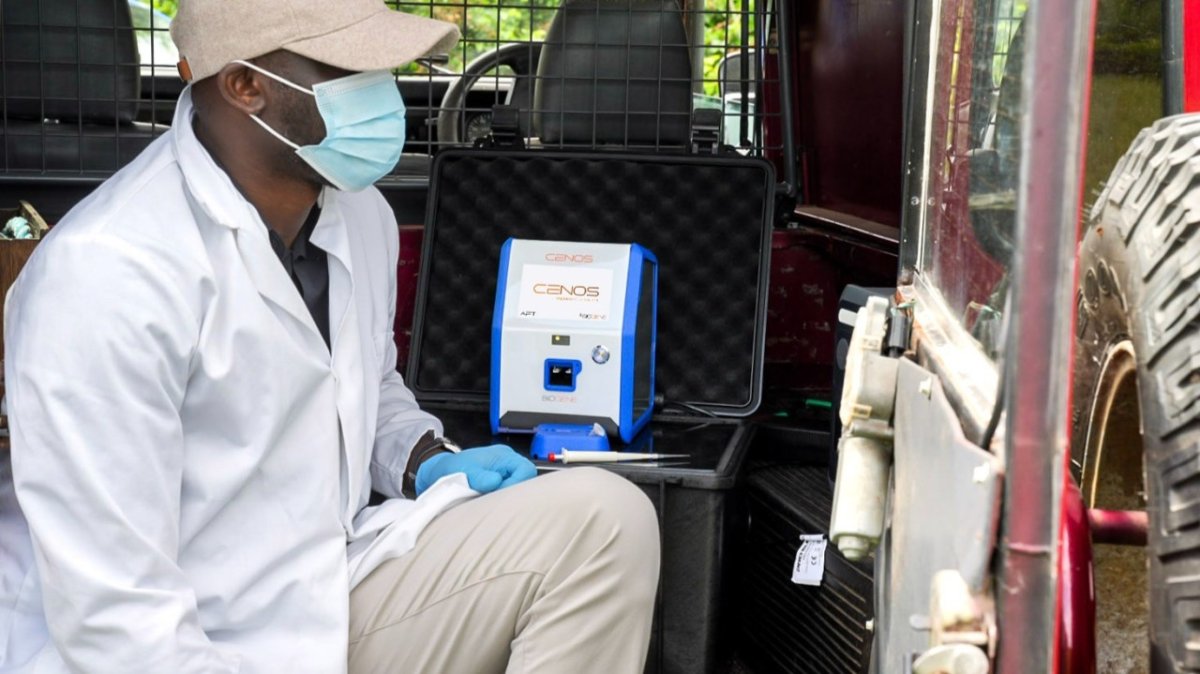
Why it matters for One Health, outbreak control, and real-time decisions
- Blood is the clinically critical matrix for several high-consequence infections because viraemia or systemic spread appears early, while the decision window stays short.
- That is true in both human and animal health. Whole blood is a priority specimen for Ebola testing, and EDTA-blood is a recommended sample for early ASFV PCR detection.
- The gap is no longer only analytical sensitivity. The gap is how fast a result can move from sample to action without transport, extraction, and laboratory dependence.
High-Consequence Infectious Diseases
In high-consequence infections, the analytical question is not enough on its own. The operational question is whether the result appears early enough, and close enough to the patient or animal, to change the next decision. In haemorrhagic fevers such as Ebola and Marburg, whole blood or EDTA-blood sits at the centre of formal diagnostic guidance. The World Health Organization records that the West Africa Ebola outbreak infected more than 28,600 people and caused 11,325 deaths, while WHO also notes that Marburg virus disease has reached case fatality ratios as high as 88 percent in past outbreaks. CDC guidance for Ebola testing identifies whole blood preserved with EDTA as the preferred specimen. Together, these points make the blood sample operationally decisive, not just analytically useful. [1][2][3][4]
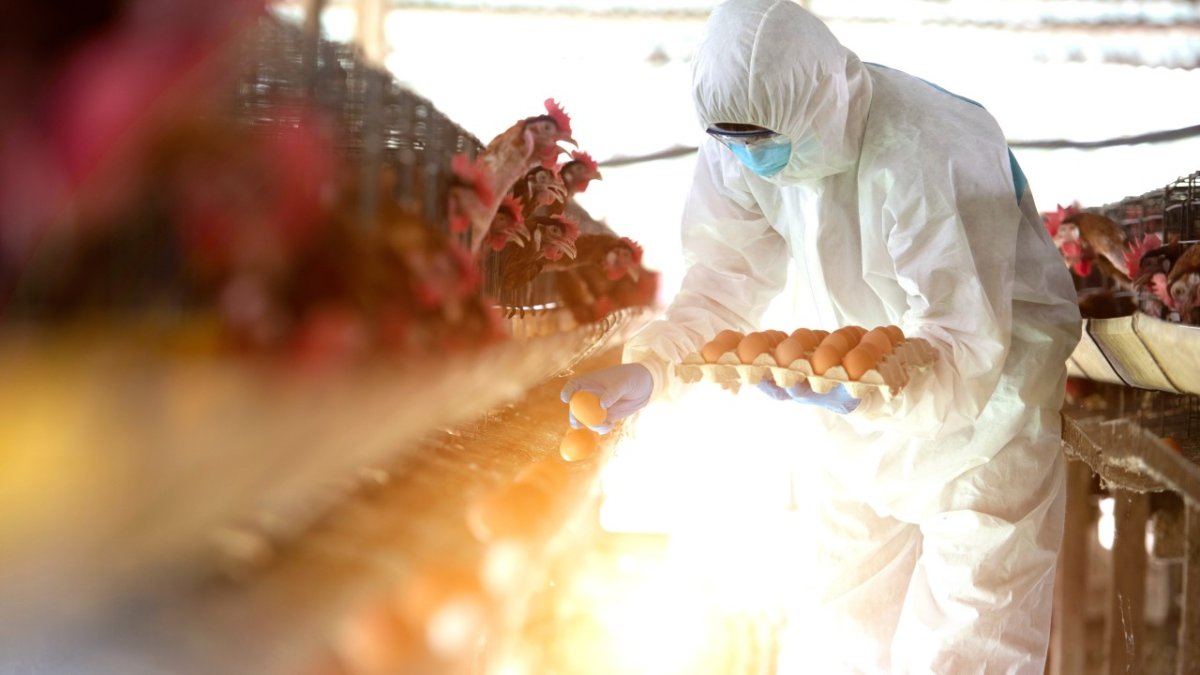
The same logic applies to animal health. WOAH guidance states that PCR can detect ASFV DNA from a very early stage of infection in EDTA-blood and serum, and reference laboratory material describes whole blood from live pigs as a preferred sample type for genome detection. For a true point-of-decision molecular system, that matters. If blood is the decisive matrix but the workflow still depends on extraction and central laboratory handling, the result arrives later than the decision window allows. [5][6]
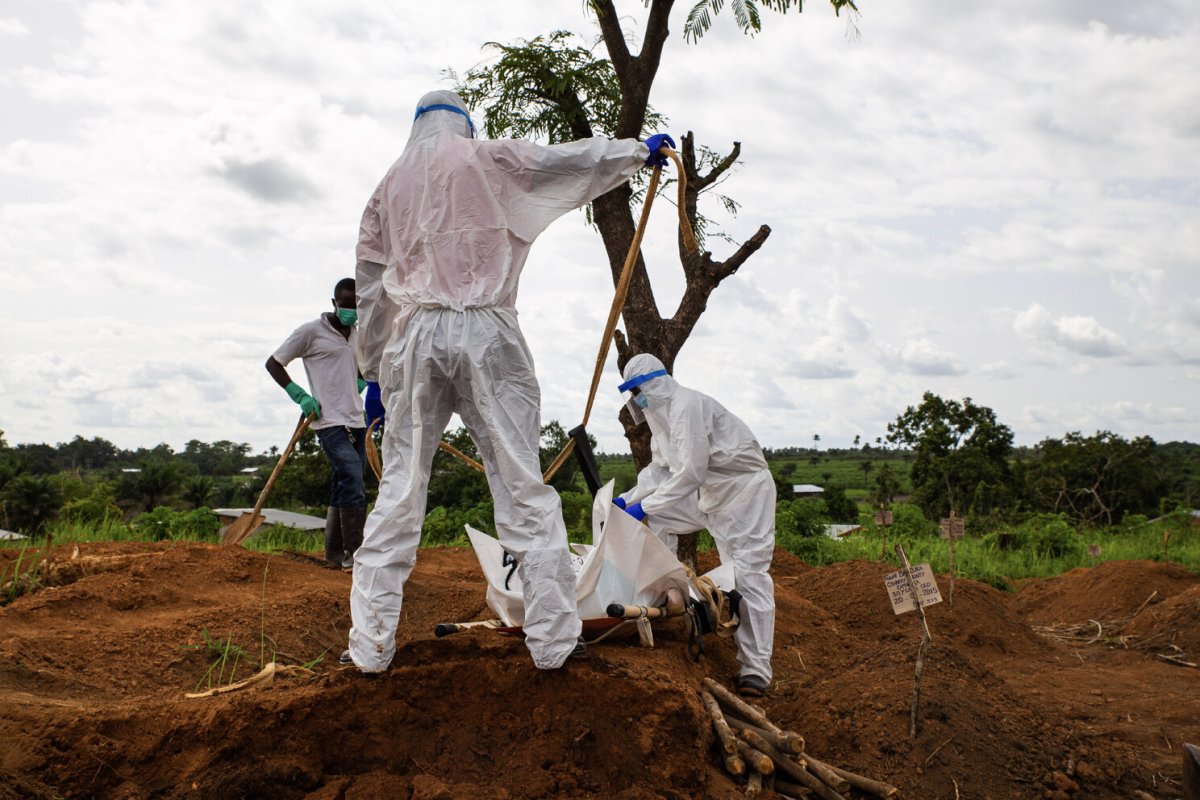

African Swine Fever Virus (ASFV)
Work conducted with The Pirbright Institute as part of an Innovate UK–funded programme demonstrates that the CENOS platform, direct-from-blood molecular detection of African Swine Fever Virus (ASFV) is both feasible and robust under controlled conditions. Across a panel of 18 ASFV strains spanning multiple genotypes, the assay showed full concordance with standard laboratory qPCR methods, while maintaining comparable Ct value trends. Importantly, performance remained stable in the presence of crude blood, as a sample input, minimal inhibition observed and reliable detection achieved with up to 16% whole blood input. Sensitivity reached detection limits of approximately 12 copies of target DNA, aligning with benchmark assays. When deployed on the CENOS platform, results were generated approximately four times faster than conventional workflows, while detecting all strains and dilutions identified by the reference lab method with sample extraction carried out.
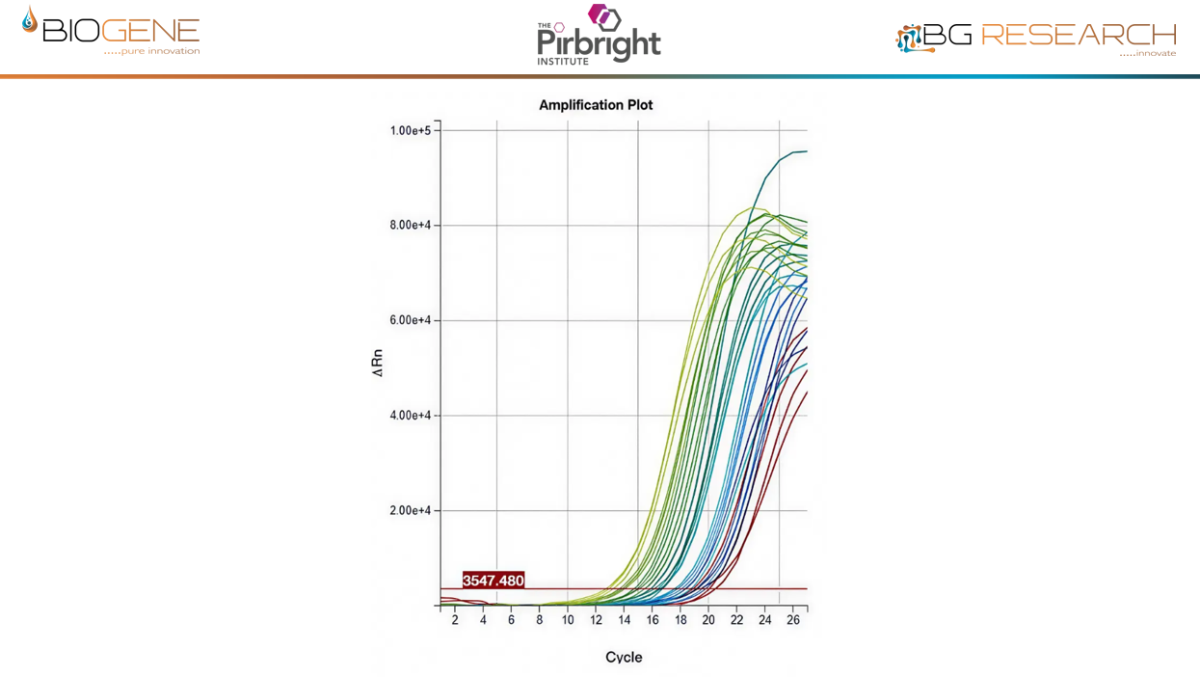
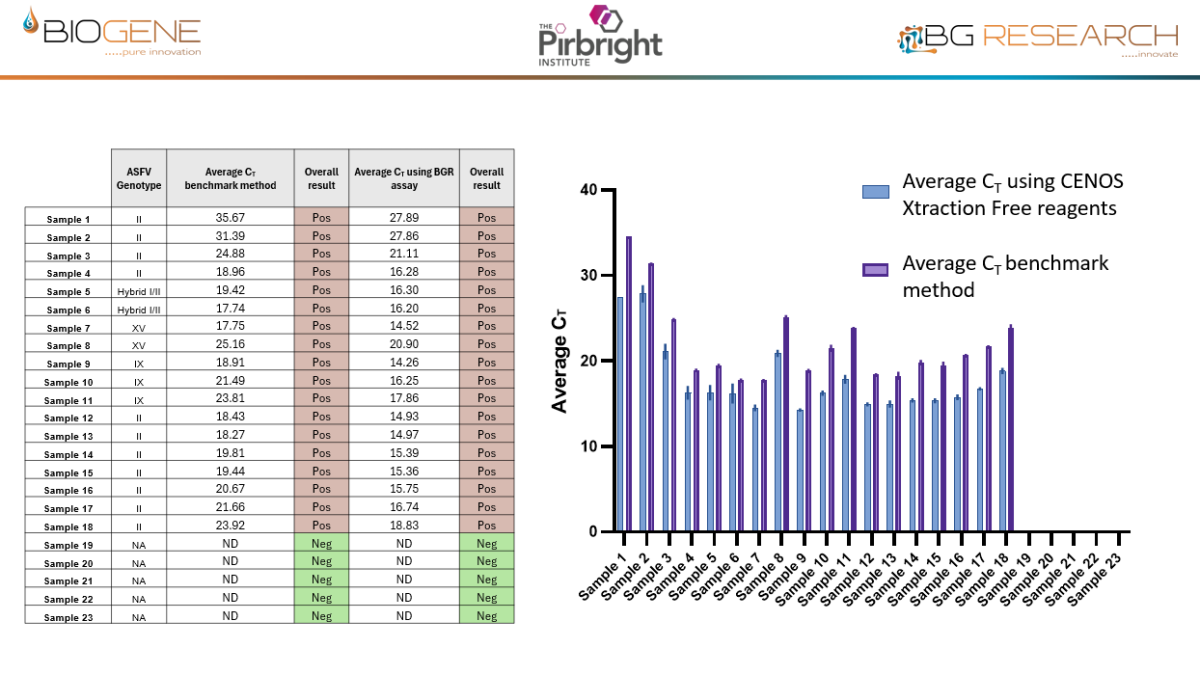
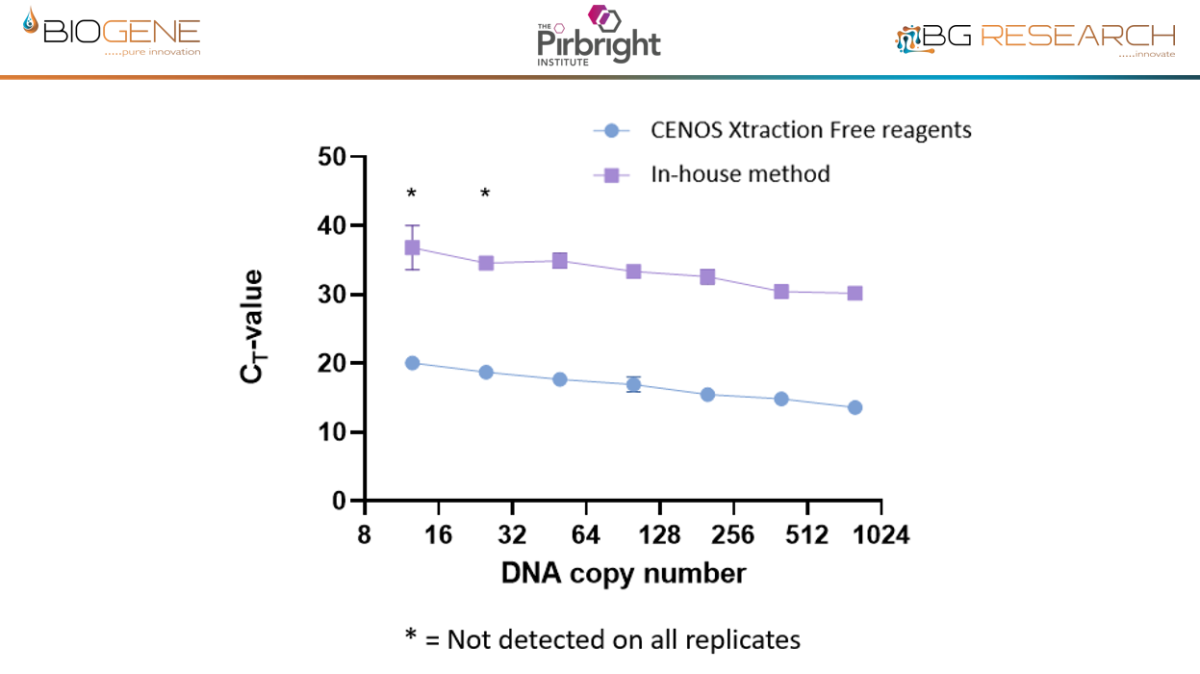
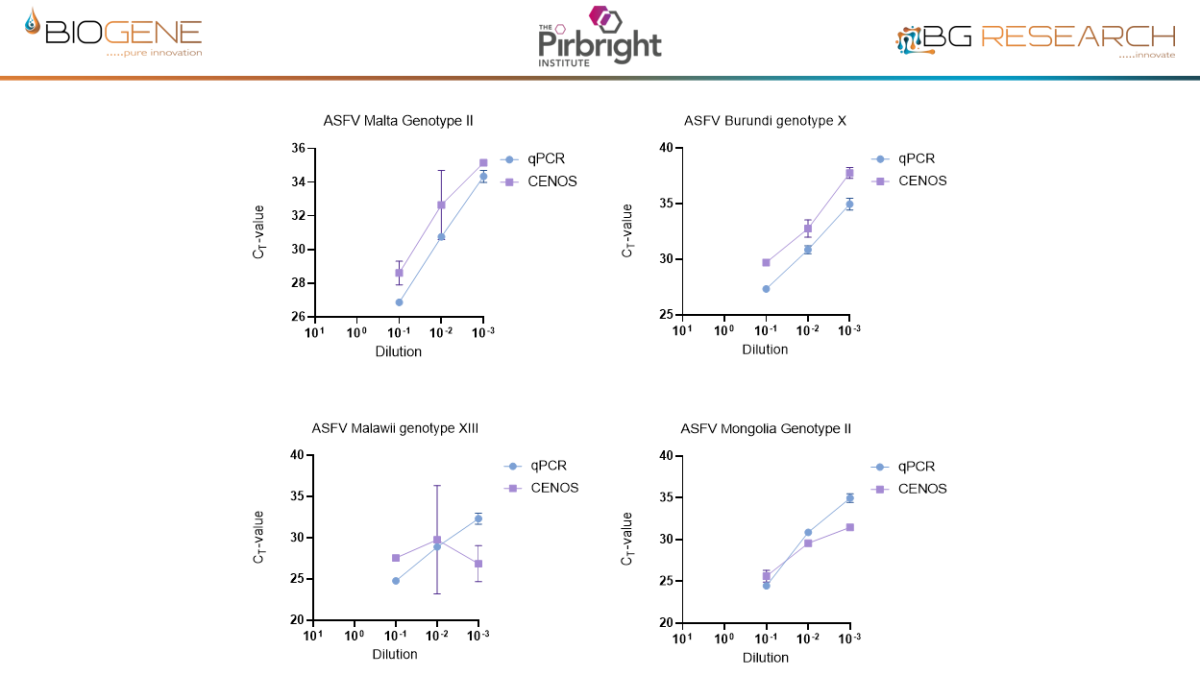
Disclaimer: Performance data generated through an Innovate UK funded research project, involving The Pirbright Institute, demonstrate the rapid molecular detection capability of the CENOS platform under defined study conditions. The participating institutions do not endorse the product
ASFV shows the cost of delay with unusual clarity
African swine fever is one of the clearest examples of why timing has economic value. FAO guidance describes ASF as a severe haemorrhagic disease of pigs with lethality up to 100 percent for highly virulent strains. Separate FAO material describes the disease as a serious transboundary threat with major consequences for food security and livelihoods. In China, a Nature Food analysis estimated total losses equal to 0.78 percent of GDP in 2019, with scenario analyses showing even larger shocks under worse production or price assumptions. [9][10][11]
This remains a current, not historical, problem. WOAH reported in early 2026 that, since January 2022, 71 countries and territories had reported ASF presence, with more than 1.1 million pig cases and more than 2.3 million animal losses in domestic pigs. DEFRA's February 2026 outbreak assessment also reported that ASF continued in domestic pigs and wild boar across Europe. These figures underline the central point: in diseases like ASFV, delay creates biological spread first and economic damage immediately after. [12][13]

Why current molecular workflows still miss the decision window
Standard molecular testing remains powerful, but the workflow still carries structural friction. CDC Ebola guidance specifies whole-blood collection, packaging, and shipment requirements, while the diagnostic literature continues to treat nucleic-acid extraction as a normal part of the RT-PCR pathway. Those steps protect analytical performance, but they also lock speed to transport, trained handling, and laboratory infrastructure. Even when assay sensitivity is strong, the pathway can still be too slow for field outbreak control, penside decisions, border screening, or urgent triage. [3][14]

One Health pressure keeps increasing
The One Health case is already data-rich. WHO describes zoonoses as a major public-health problem and states that around 60 percent of emerging infectious diseases are zoonotic. WHO regional material adds that zoonoses cause about one billion cases of illness and millions of deaths each year. This matters because surveillance, triage, outbreak control, and trade decisions do not happen in separate worlds. Human health, animal health, and environmental exposure interact continuously, which raises the value of molecular tools that can operate at the decision point rather than only at the reference laboratory. [7][8]
The Global Early Warning System (GLEWS+) shows how effectively we can connect intelligence across human and animal health. The remaining challenge is speed of response — turning that insight into actionable diagnostics at the point of decision: https://www.glews.net/

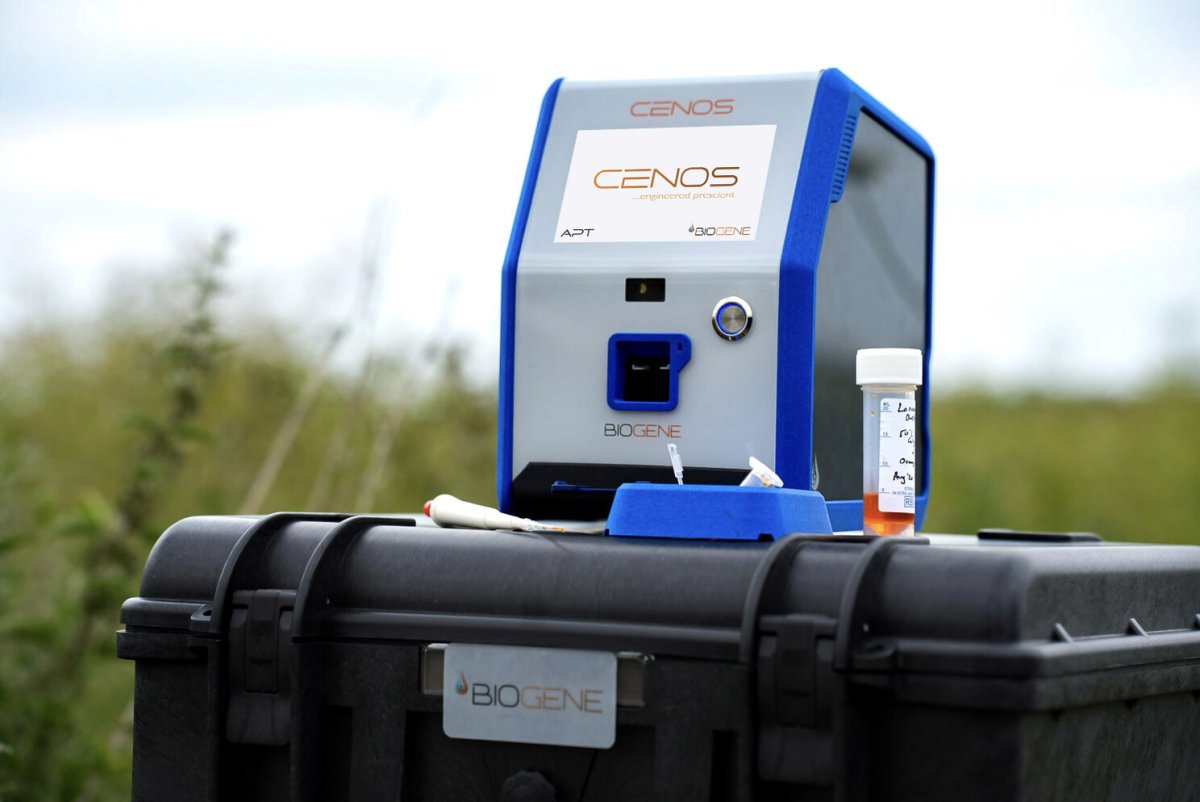
What changes with direct-from-blood detection using CENOS
The strategic value of direct-from-blood testing is simple: it removes the extraction bottleneck from the time-critical part of the workflow. That does not make blood less challenging as a matrix; it makes the platform more useful if it can handle that matrix directly. For CENOS, the claim is not that blood has suddenly become easy. The claim is that blood is the matrix that matters most in several high consequence use cases, and that a platform able to test directly from blood has the potential to move molecular diagnostics from reporting toward action.
That is why the ASFV work with The Pirbright Institute matters so much in this narrative. It gives a concrete and credible pointer that direct-from-blood molecular testing can align with reference laboratory expectations in a real veterinary high-consequence setting, while opening a route to faster and more deployable response.

References
[1] World Health Organization. Ebola outbreak 2014-2016 - West Africa. WHO.Direct detection of ASFV, spiked into whole pig blood from 6-800 virions/rxn. LLOD is <1,000 virions/ml or eluate.Find out more
[2] World Health Organization. Marburg virus disease. WHO fact sheet, updated 20 January 2025.
[3] Centers for Disease Control and Prevention. Guidance for Collection, Transport and Submission of Specimens for Ebola Virus Testing in the United States.
[4] CDC Ebola Virus NP Real-time RT-PCR Assay: Fact Sheet for Health Care Providers.
[5] World Organisation for Animal Health. African swine fever (ASF), Terrestrial Manual chapter.
[6] Cafariello C. et al. Spleen swabs for sensitive and high-throughput detection of African swine fever virus. 2024. Summary notes whole blood from live pigs as a preferred WOAH-recommended sample type for real-time PCR.
[7] World Health Organization. Zoonoses fact sheet.
[8] World Health Organization, Eastern Mediterranean Regional Office. Zoonotic disease: emerging public health threats in the Region.
[9] FAO. African swine fever: detection and diagnosis - a manual for veterinarians.
[10] FAO. Recognizing African swine fever: a field manual.
[11] You S. et al. African swine fever outbreaks in China led to gross domestic product and economic losses. Nature Food, 2021.
[12] World Organisation for Animal Health. ASF situation report, early 2026 update.
[13] Department for Environment, Food and Rural Affairs. African swine fever in Europe - updated outbreak assessment, 24 February 2026.
[14] Bettini A. et al. Diagnostics of Ebola virus. 2023 review noting extraction as the standard RNA preparation step for RT-PCR.


