Global Context
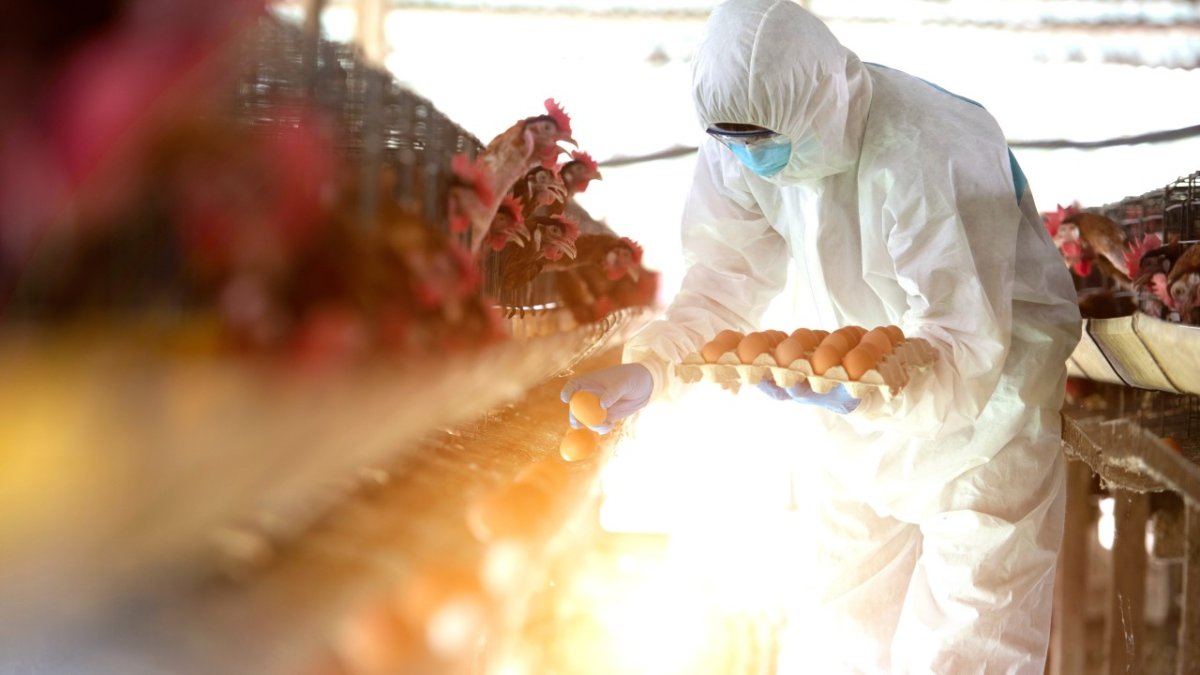
Highly pathogenic avian influenza (HPAIV) H5N1 has expanded dramatically in geographic range in recent years.
Since 2020 outbreaks have been reported in more than 80 countries across Europe, Asia, Africa and the Americas.
Control programmes and preventive culling have resulted in the loss of more than 500 million poultry globally.
The 2022–2024 outbreak waves represented the largest recorded epizootic of avian influenza in several regions.
Wild bird mortality events have been documented across Europe and the Americas, highlighting the role of migratory species in virus dissemination.
Spillover events into mammals have also been recorded, including:
- foxes
- seals
- sea lions
- farmed mink
These events have intensified monitoring efforts because of the zoonotic potential of influenza viruses.

Newcastle Disease
Alongside avian influenza, Newcastle disease virus (NDV) remains one of the most economically damaging poultry pathogens worldwide.
In unvaccinated flocks mortality may reach 90–100 percent.
In many regions where veterinary infrastructure is limited, NDV outbreaks occur regularly and can cause greater cumulative economic losses than avian influenza because the virus frequently circulates endemically.
Limitations of Conventional Diagnostics
Diagnostic confirmation for notifiable avian diseases typically relies on laboratory real-time PCR testing.
These workflows require:
- sample transport to specialised laboratories
- specialist equipment
- nucleic-acid extraction procedures
- trained molecular diagnostic personnel
While highly sensitive and specific, these requirements introduce delays during outbreaks and limit diagnostic capacity in areas lacking laboratory infrastructure.
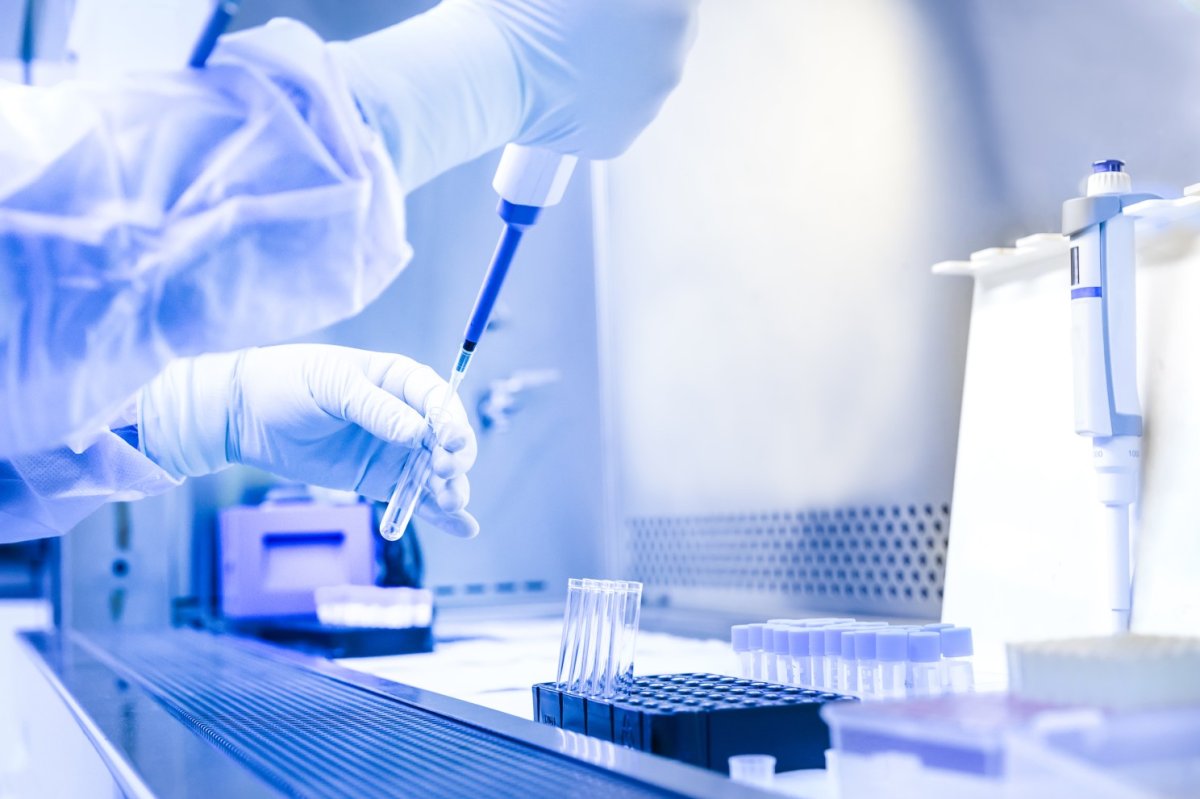

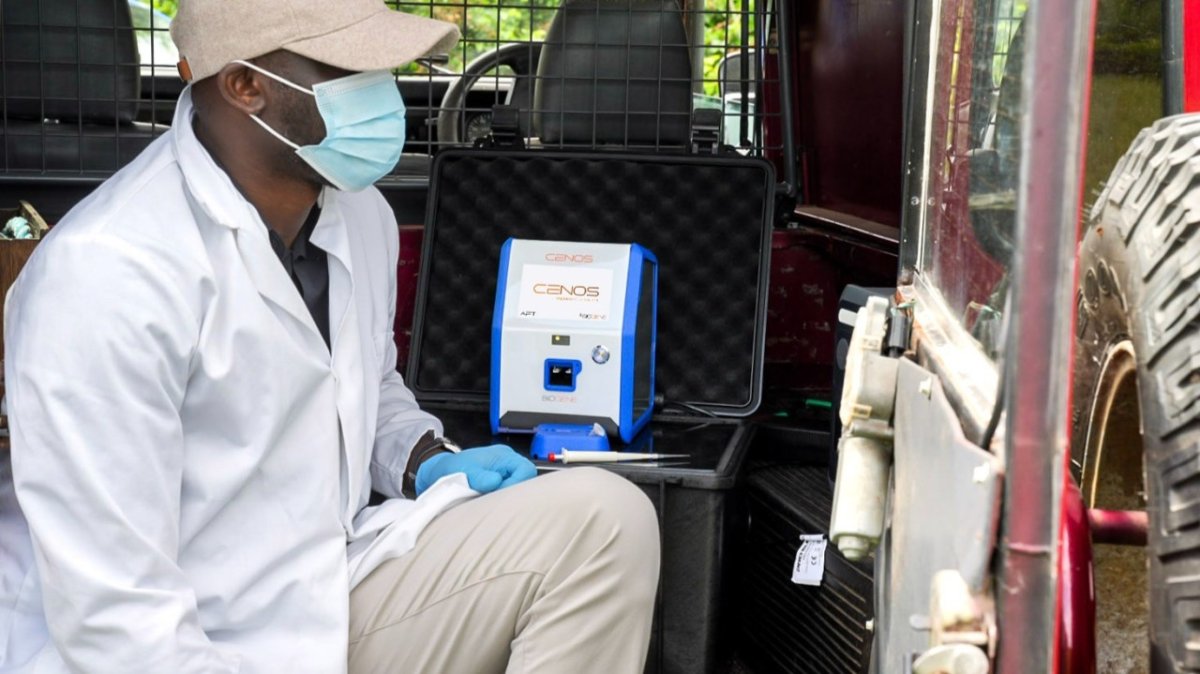
Direct Molecular Detection using CENOS
Research collaboration between BioGene and the Animal and Plant Health Agency (APHA) evaluated a diagnostic approach using the CENOS molecular platform.
The platform enables real-time RT-PCR detection directly from crude biological samples, including:
- cloacal swabs
- oropharyngeal swabs
Viral particles are lysed within the reaction environment, RNA stabilised and reverse transcription and amplification occur within a single reaction workflow for each target.
This extraction-free approach simplifies the diagnostic process while maintaining analytical sensitivity.
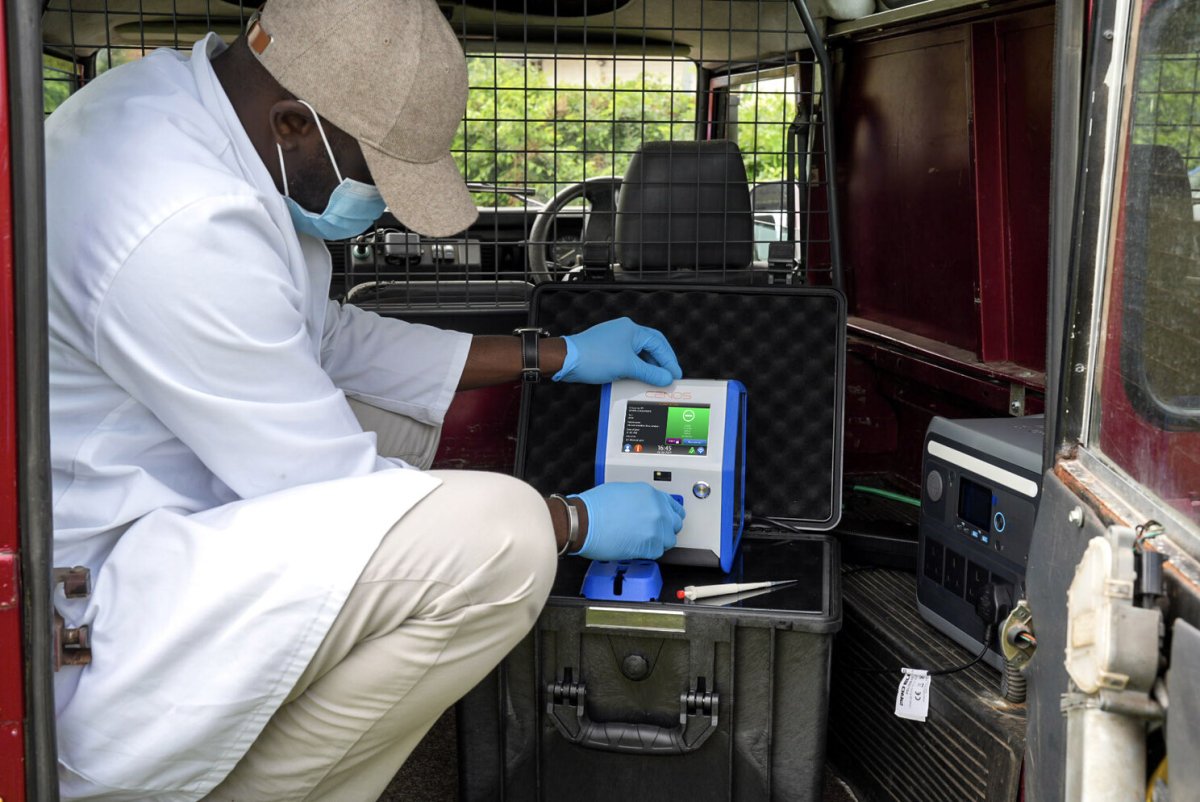

Validation results
Evaluation within APHA high-containment laboratories compared the novel assays with accredited frontline PCR diagnostics.
Key findings
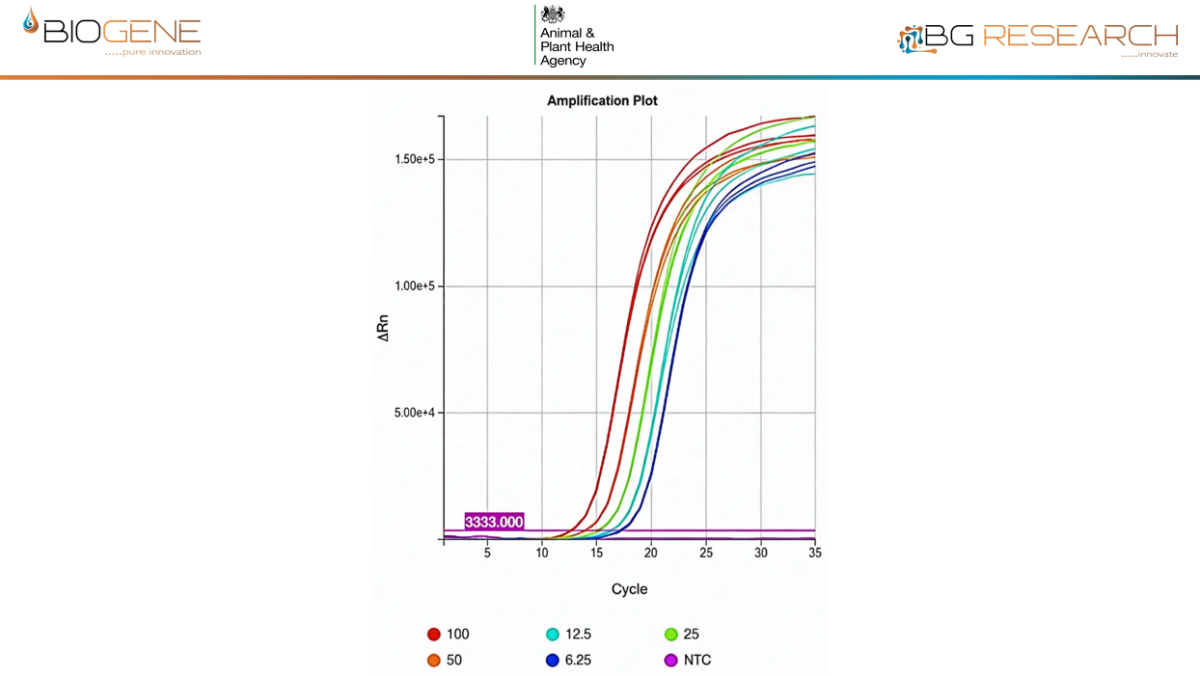
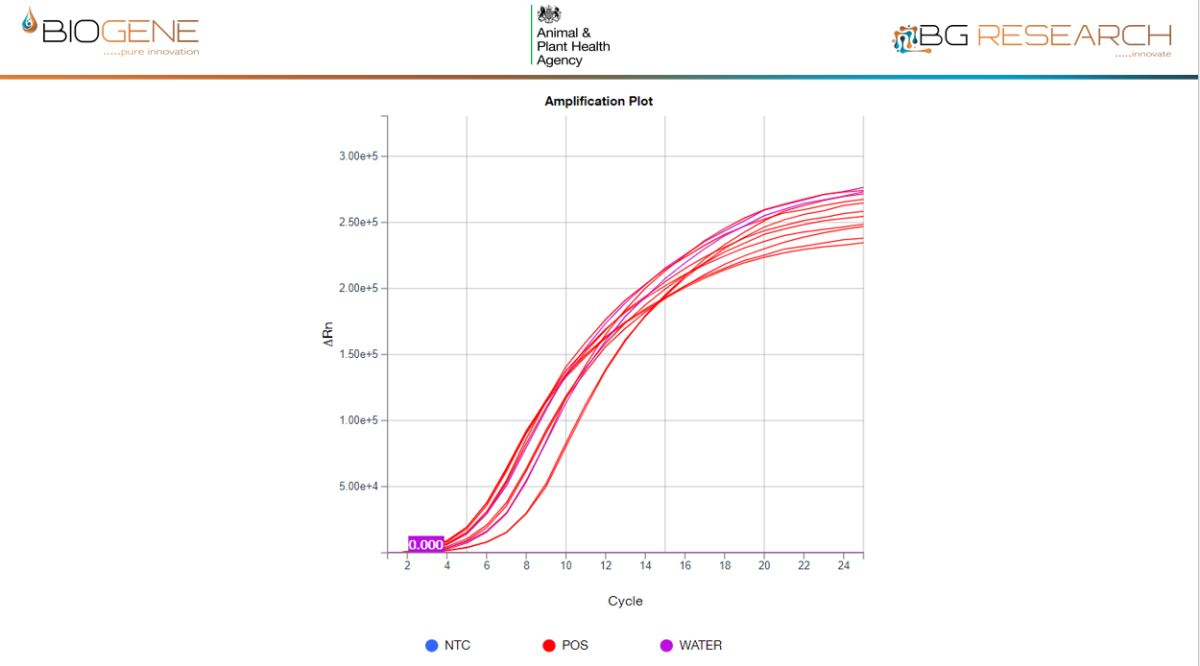
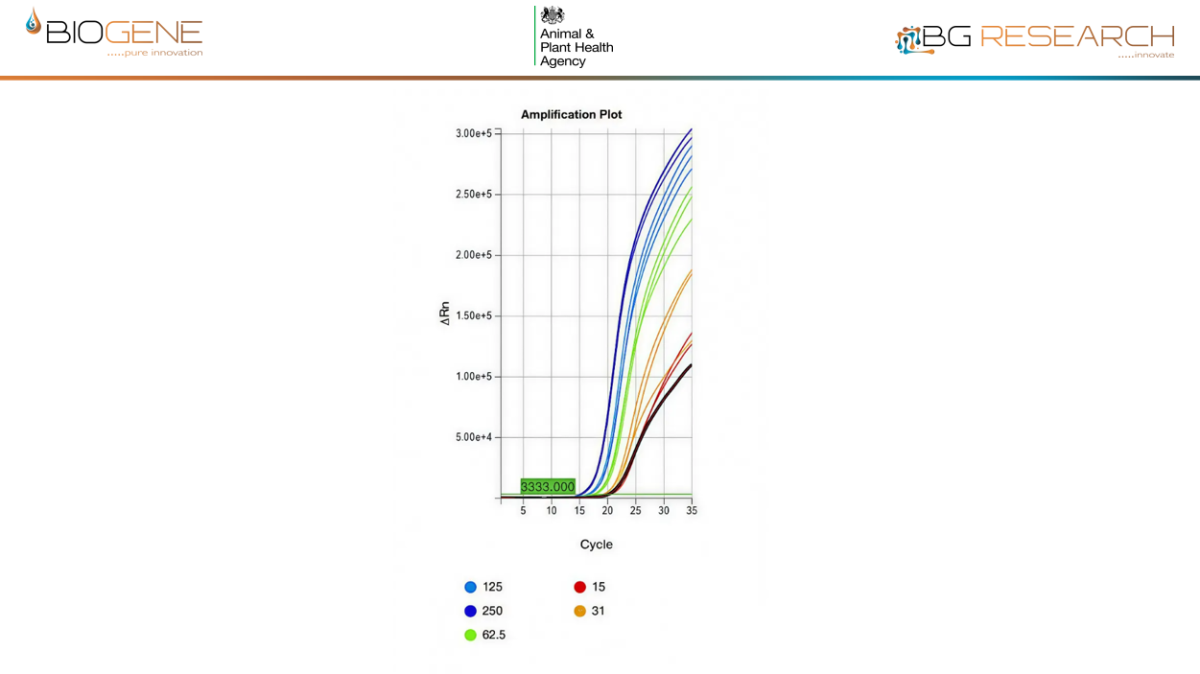
- Predicted detection of >99.9% of circulating AIV and NDV genomic sequences
- Successful detection across multiple viral genotypes
- Sensitivity and specificity comparable to reference PCR assays
- Reliable detection from cloacal and oropharyngeal swabs without nucleic-acid extraction
AIV Data
NDV Data
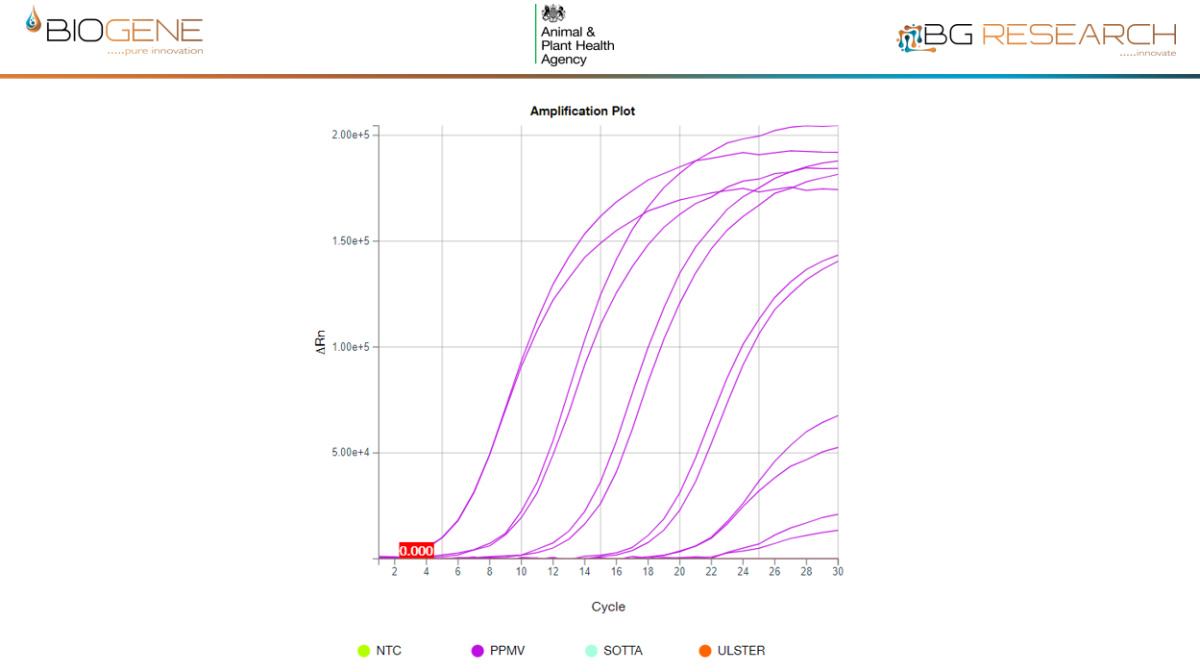
Disclaimer: performance data generated through an Innovate UK funded research project, involving the Animal & Plant Health Agency (APHA), demonstrate the rapid molecular detection capability of the CENOS platform under defined study conditions. The participating institutions do not endorse the product.
Implications for surveillance
Extraction-free molecular diagnostics could significantly improve outbreak response and surveillance capacity.
Testing closer to farms, wildlife monitoring sites or border inspection points enables:
- faster detection at the source of outbreak
- streamlined workflows
- reduced diagnostic costs
- expanded surveillance coverage
Such decentralised diagnostic capability may play an important role in strengthening global One Health surveillance systems.
Global Data
- >500 million poultry affected globally by avian influenza since 2020 (WOAH)
- HPAIV H5N1 detected in more than 80 countries
- Major wildlife mortality events reported across Europe and the Americas
- Increasing spillover documented in mammals including foxes, mink, seals and sea lions
- >880 confirmed human H5N1 infections recorded since 2003 (WHO)
- Newcastle disease mortality may reach 90–100% in susceptible poultry flocks

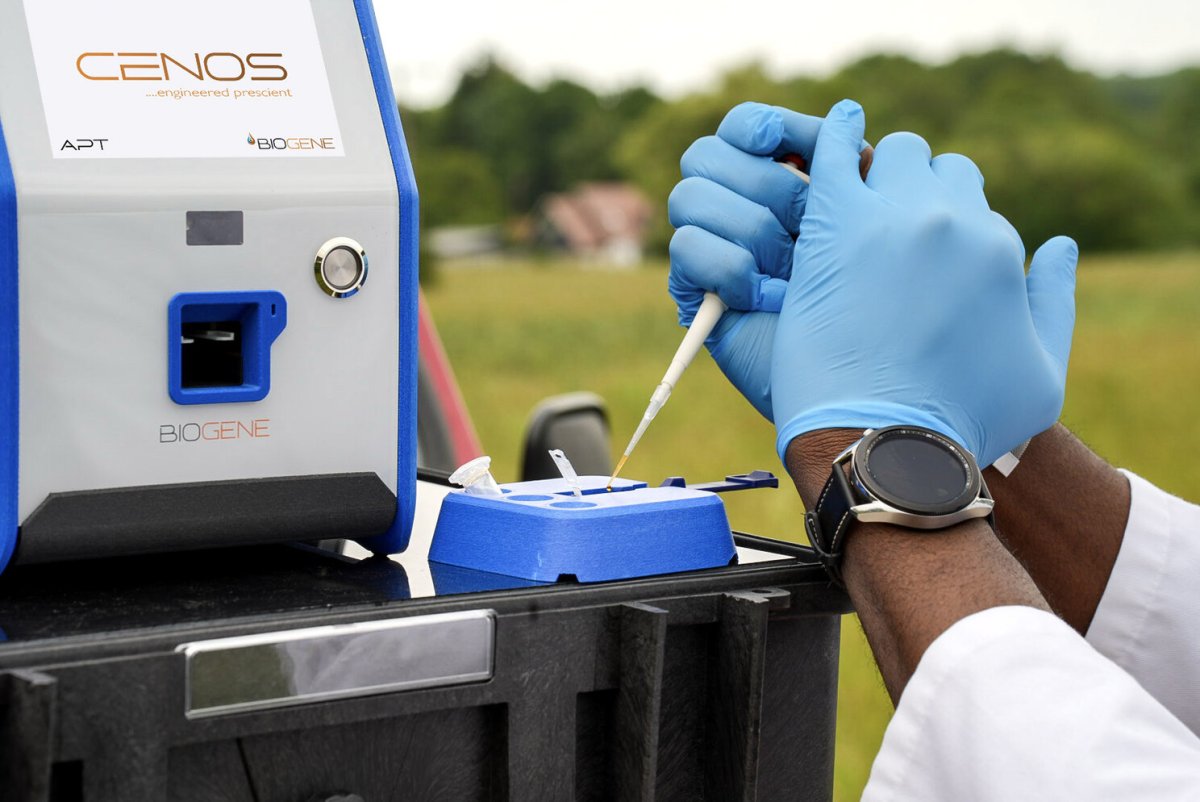
Technology Overview – CENOS Extraction-Free Workflow
- Direct crude sample input (cloacal or oropharyngeal swab)
- Heat and reagent chemistry lyse viral particles
- Viral RNA stabilised within the reaction mixture
- Reverse transcription converts RNA into cDNA
- Real-time PCR amplification detects viral genome sequences (one assay per target)
This workflow eliminates nucleic-acid extraction while maintaining PCR-level analytical performance.

References
World Organisation for Animal Health (WOAH) — Global situation of avian influenza
Food and Agriculture Organization (FAO) — Global avian influenza updates
European Food Safety Authority (EFSA) — Avian influenza surveillance reports
World Health Organization (WHO) — Human infections with avian influenza A(H5N1)
Animal and Plant Health Agency (APHA) — Reference laboratory for avian influenza and Newcastle disease